The growing polypeptide connected to the tRNA in the P site is detached from the tRNA in the P site and a peptide bond is formed between the last amino acids of the polypeptide and the amino acid still attached to the tRNA in the A site. Now the P site contains the beginning of the peptide chain of the protein to be encoded and the A site has the next amino acid to be added to the peptide chain. For fast and accurate recognition of the appropriate tRNA, the ribosome utilizes large conformational changes ( conformational proofreading). This binding is facilitated by elongation factor-Tu (EF-Tu), a small GTPase. Įlongation starts when the fMet-tRNA enters the P site, causing a conformational change which opens the A site for the new aminoacyl-tRNA to bind. The growing protein exits the ribosome through the polypeptide exit tunnel in the large subunit. Elongation Įlongation of the polypeptide chain involves addition of amino acids to the carboxyl end of the growing chain. Leaderless initiation can occur when the complete 70S ribosome binds initiation factors and fMet-tRNAs, but on mRNAs that lack 5' UTRs and have a start codon on their 3' end. 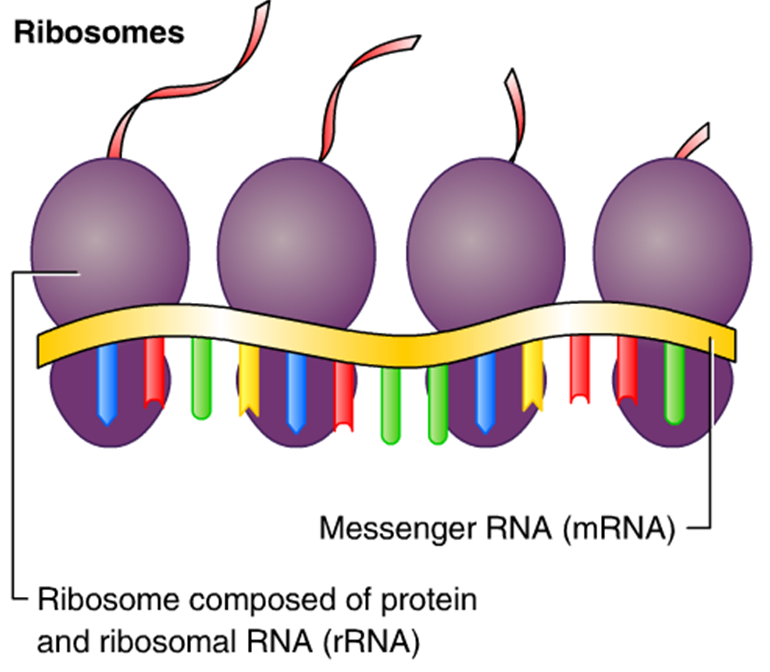
This mode is thought to be important for the translation of genes that are clustered in poly-cistronic operons, where the canonical binding mode can be disruptive due to small distances between neighboring genes on the same mRNA molecule. In the scanning mode, the complete 70S ribosome while already on mRNA, can bind to initiation factors fMet-tRNA and initiate translation by scanning the mRNA for a start site. In the canonical mode, the 30S ribosomal subunit, initiation factors, and initiator fMet-tRNA bind to the mRNA to form the pre-initiation complex, which then recruits the 50S ribosomal subunit to start translation elongation. There are three modes of initiation: the canonical 30S binding model, the 70S scanning mode, and leaderless initiation. Two studies have independently shown that 17 or more non-AUG start codons may initiate translation in E. Well-known coding regions that do not have AUG initiation codons are those of lacI (GUG) and lacA (UUG) in the E. During the formation of the initiation complex, these complementary nucleotide sequences pair to form a double stranded RNA structure that binds the mRNA to the ribosome in such a way that the initiation codon is placed at the P site. This sequence has been evolutionarily conserved and plays a major role in the microbial world we know today. The Shine-Dalgarno sequence is complementary to a pyrimidine rich region on the 16S rRNA component of the 30S subunit. The 30S subunit binds to the mRNA template at a purine-rich region (the Shine-Dalgarno sequence) upstream of the AUG initiation codon. The selection of an initiation site (usually an AUG codon) depends on the interaction between the 30S subunit and the mRNA template. And the E site which is the exit site of the now uncharged tRNA after it gives its amino acid to the growing peptide chain. The P site is where the peptidyl tRNA is formed in the ribosome. The A site is the point of entry for the aminoacyl tRNA (except for the first aminoacyl tRNA, which enters at the P site). The ribosome has three active sites: the A site, the P site, and the E site. Variations in the mechanism can be anticipated. Initiation of translation in bacteria involves the assembly of the components of the translation system, which are: the two ribosomal subunits ( 50S and 30S subunits) the mature mRNA to be translated the tRNA charged with N-formylmethionine (the first amino acid in the nascent peptide) guanosine triphosphate (GTP) as a source of energy, and the three prokaryotic initiation factors IF1, IF2, and IF3, which help the assembly of the initiation complex. Bacterial translation is the process by which messenger RNA is translated into proteins in bacteria.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
